|
pKiss is a tool for folding RNA secondary structures, including two limited classes of pseudoknots.
As pKiss is the successor of pknotsRG, the first pseudoknot class is the canonical simple recursive pseudoknot from pknotsRG. The new class are canonical simple recursive kissing hairpins.
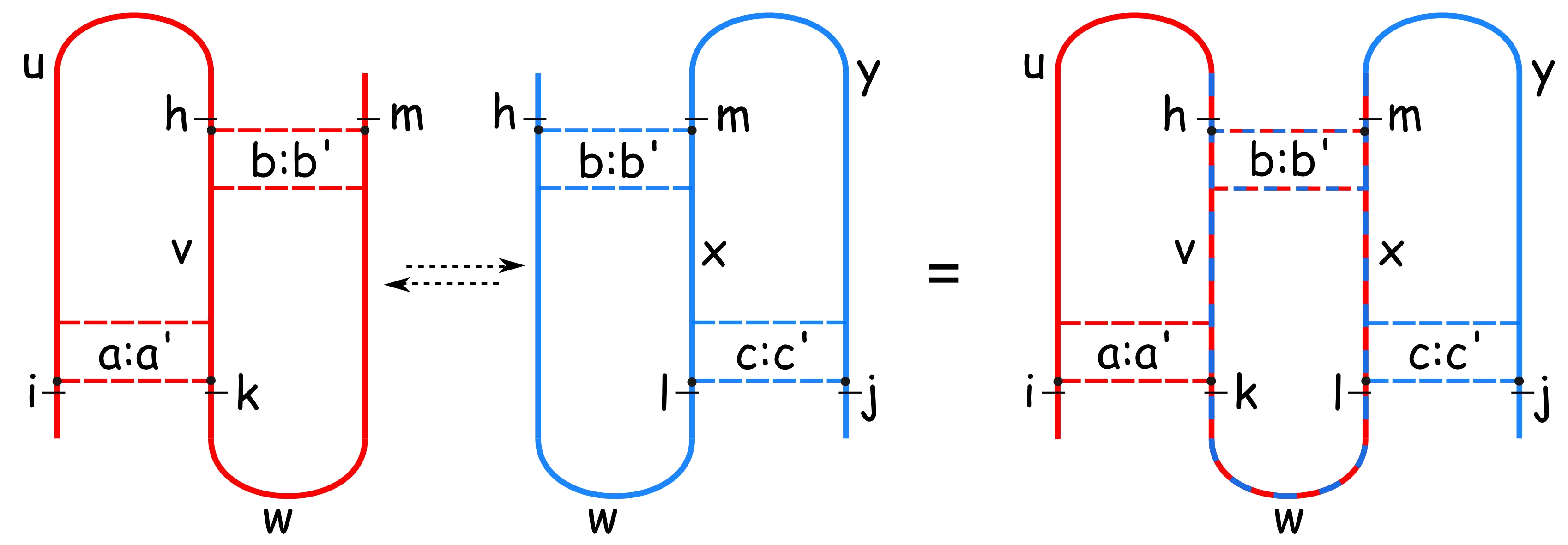
To retain the fast runtime while adding the more complicated kissing hairpin class, we heuristically construct a kissing hairpin from an overlay of two canonical simple recursive pseudoknots. By applying this trick, we loose a bit thoroughness. Due to our evaluations, performance is still very good.
Should you feel uncomfortable with this situation, pKiss offers three more strategies to compute kissing hairpins. These strategies are either slower or consume more memory.
We strongly suggest using Strategy A. See the manual for details.
| Strategy |
time complexity |
space complexity |
| Strategy A |
O(n4) |
O(n2) |
| Strategy B |
O(n4) |
O(n3) |
| Strategy C |
O(n5) |
O(n2) |
| Strategy D |
O(n6) |
O(n2) |
Author: S. Janssen
built on June 25 2015 (32:49335b54b1ba)
|
